2026 (Bolded denotes Phillips lab members)
Tharp CR, Catalano C, Khalifeh A, Ghaffari-Kashani S, Huang R, Kang G, Scapin G, Phillips AM.
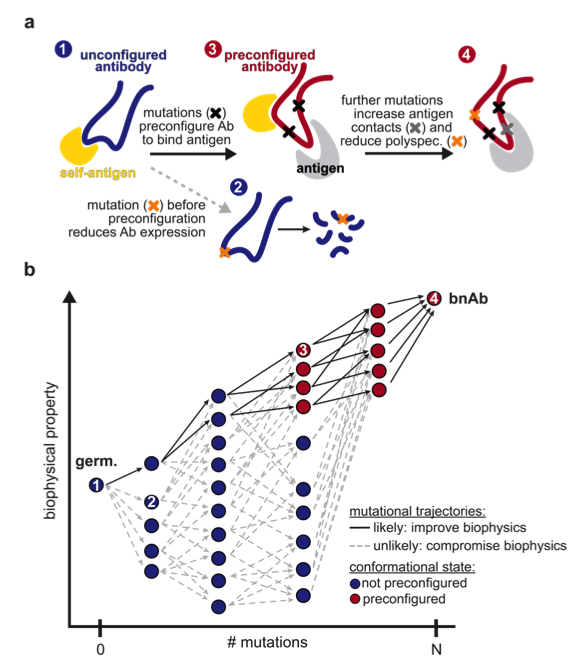
Biophysical trade-offs in antibody evolution are resolved by conformation-mediated epistasis.
 BioRxiv. 2026.
BioRxiv. 2026.
Angela’s publications as a postdoc
Dupic T*, Phillips AM*, Desai MM. Protein sequence landscapes are not so simple: on reference-free versus reference-based inference. BioRxiv. 2024.
Phillips AM*†, Maurer DP*, Brooks CR, Dupic T, Schmidt AG, Desai MM†. Hierarchical sequence-affinity landscapes shape the evolution of breadth in an anti-influenza receptor binding site antibody. eLife. 2023; 12:e83628.
Moulana A*, Dupic T*, Phillips AM*†, Chang J*, Nieves S, Roffler AA, Starr TN, Greaney AJ, Bloom JD, Desai MM†. The landscape of antibody binding affinity in SARS-CoV-2 Omicron BA.1 evolution. eLife. 2023; 12:e83442.
Moulana A, Dupic T, Phillips AM, Desai MM. Genotype-phenotype landscapes for immune-pathogen coevolution. Trends Immunol. 2023;44(5):384-96.
Moulana A*, Dupic T*, Phillips AM*†, Chang J*, Nieves S, Roffler AA, Starr TN, Greaney AJ, Bloom JD, Desai MM†. Compensatory epistasis minimizes trade-offs between antibody escape and ACE2 affinity for SARS-CoV-2 Omicron BA.1. Nat Comm. 2022; 13:7011.
Phillips AM*, Lawrence KR*, Moulana A, Dupic T, Chang J, Johnson MS, Cvijovic I, Mora T, Walczak AM, Desai MM. Binding affinity landscapes constrain the evolution of broadly neutralizing anti-influenza antibodies. eLife. 2021; 10:e71393.
Bakerlee CW*, Phillips AM*, Nguyen Ba AN, Desai MM. Dynamics and variability in the pleiotropic effects of adaptation in laboratory budding yeast populations. eLife. 2021; 10:e70918.
Tung S*, Bakerlee CW*, Phillips AM, Nguyen Ba AN, Desai MM. The genetic basis of differential autodiploidization in evolving yeast populations. G3. 2021; 11.
Johnson MJ, Gopalakrishnan S, Goyal J, Dillingham ME, Bakerlee CW, Humphrey PT, Jagdish T, Jerison ER, Kosheleva K, Lawrence KR, Min J, Moulana A, Phillips AM, Piper JC, Purkanti R, Rego-Costa A, McDonald MJ, Nguyen Ba AN, Desai MM. Phenotypic and molecular evolution across 10,000 generations in laboratory budding yeast populations. eLife. 2021; 10:e63910.
Angela’s publications as a graduate student
Phillips AM, Doud MB, Gonzalez LO, Butty VL, Lin Y-S, Bloom JD, Shoulders MD. Enhanced ER proteostasis and temperature differentially impact the mutational tolerance of influenza hemagglutinin. eLife. 2018; 7:e38795.
Phillips AM*, Ponomarenko AI*, Chen K, Ashenberg O, Miao J, McHugh SM, Butty VL, Whittaker CA, Moore CL, Bloom JD, Lin Y-S, Shoulders MD. Destabilized adaptive influenza variants critical for innate immune system escape are potentiated by host chaperones. PLoS Biology. 2018; 16:e3000008.
Phillips AM, Gonzalez LO, Nekongo EE, Ponomarenko AI, McHugh SM, Butty VL, Levine SS, Lin Y-S, Mirny LA, Shoulders MD. Influenza evolution is modulated by host proteostasis mechanisms. eLife. 2017; 6:e28652.
Phillips AM, Shoulders MD. Commentary: The path of least resistance: mechanisms to reduce influenza’s sensitivity to oseltamivir. Journal of Molecular Biology. 2016; 428:533–537.
Yoon, J.*, Nekongo, E.E.*, …Phillips A.M. et al. The endoplasmic reticulum proteostasis network profoundly shapes the protein sequence space accessible to HIV envelope. PloS Biology. 2022; 20:e3001569.
For a full list of Angela’s publications, see her Google Scholar profile